---
title: "Introduction to KEGGlincs"
author: "Shana White"
date: "03/23/2017"
---
by [Shana White](https://github.com/shanawhite-UC)
Installation:
```{r}
#Make sure that the following bioconductor packages are installed
if (!requireNamespace("BiocManager", quietly=TRUE))
install.packages("BiocManager")
BiocManager::install(c("hgu133a.db", "KEGGgraph", "KEGGREST", "KOdata"))
#Download package
BiocManager::install("KEGGlincs")
#Load/activate package for use
library(KEGGlincs)
```
# Overview of package

----
KEGGlincs is an R package that provides a seamless interface for viewing detailed versions of KEGG pathways in Cytoscape such that the exact relationships (*edges*) that exist between pathway elements (*nodes*) are visualized in a meaningful manner.
This package is intended to be used as a tool to accurately regenerate KEGG pathways with *expanded* edges that are marked-up (in terms of color or color+width) to portray specific *edge attributes*. Specifically,
- the edges are *expanded* to represent all relationships between genes encoded by the [KGML file](http://www.kegg.jp/kegg/xml/) but not necessarily represented in the original [KEGG] pathway. This 'expansion' occurs either because:
- one node may represent many [often paralogous] genes.
- a node is of type 'group' and represents two or more genes that are part of the same functional complex.
- the *edge attributes* represented can be specified by the user as either:
- Functional - the exact *type* and [in some cases] *subtype* of the relationship defined by the KGML file.
- Data driven - based on quantitative experimental evidence for gene-gene relationships generated by experimental data. As an added feature and motivating example, this package gives users an opportunity to use [LINCS L1000 knock-out data](http://www.lincscloud.org/l1000/) in order to analyze/visualize the pathway edges as they pertain to specific [cell lines](http://www.lincscloud.org/cell_types/).
Aside from edge-specific annotation/mark-up, a unique feature of this package lies in the ability for users to generate graph objects in R and send them to [Cytoscape](http://www.cytoscape.org/) for an interactive visualization experience via utilities developed by the [cyREST team](https://github.com/idekerlab/cy-rest-R).
The following example showcases the ideas described above.
## Example: p53 pathway
### Original KEGG pathway:
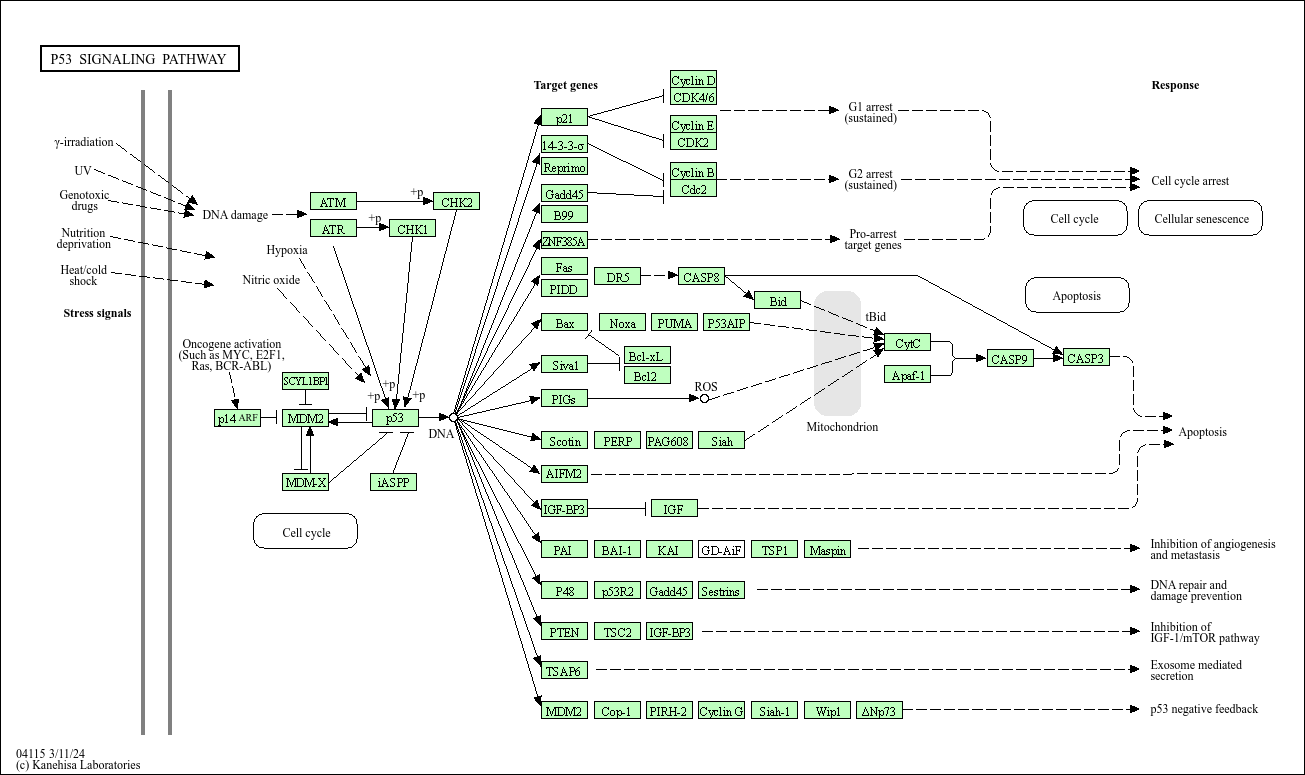
### Using KEGGlincs without adding data:
#### Detailed pathway visualized in Cytoscape
- Explicitly shows all documented edges
- Color-coded to easily interpret edge type (i.e. activation, inhibition, etc.)
- Edge color key: <a><img src="https://cdn.rawgit.com/uc-bd2k/KEGGlincs/master/vignettes/image_files/color_1.svg"/></a>
*Note: An open Cytoscape session is required for pathway visualization; i.e. make sure that Cytoscape is open before running the following commands*
```{r}
KEGG_lincs("hsa04115")
```

### Overlay pathway edges with LINCS data:
#### Pathway with edge attributes generated from LINCS L1000 knock-out data, *refined* by cell-line specific expression (default option, see below), and visualized in Cytoscape
- Explicitly shows all documented edges; edges that do not exist in the data appear but are not marked-up for interpretation [of significance].
- If baseline expression data is available for the selected cell-line, the pathway will be *refined* to only include edges between genes that are expressed in a given cell line. The nodes for unexpressed genes are colored gray.
- Widths: Magnitude of concordance for the edge in the selected cell-type
- Colors: Direction and significance for the edge in the selected cell-type (based on a modified odds-ratio score).
- Edge color key (+/- refers to *direction* of score; scores are considered significant if adjusted p-value is >= 0.05):
<a><img src="https://cdn.rawgit.com/uc-bd2k/KEGGlincs/master/vignettes/image_files/color_2.svg"/></a>
#### Pathway specific to PC3 cell-line (Prostate cancer)
```{r}
KEGG_lincs("hsa04115", "PC3")
```

#### Pathway specific to HA1E cell-line (Immortalized normal kidney epithelial)
```{r}
KEGG_lincs("hsa04115", "HA1E")
```
